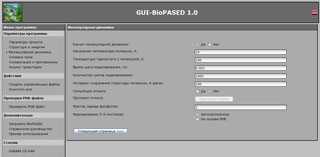
General layout of GUI-BioPASED program in a web browser window. Molecular dynamics optimization setup page.
There are several software packages that allow the user to model main types of biopolymers with molecular dynamics methods. However, the work with most of these packages is quite complicated, especially for non-experts in computational structural biology. We have created GUI-BioPASED, a web-based GUI for molecular dynamics software BioPASED, to partially automate the tasks usually responsible for the majority of errors. GUI-BioPASED creates a task as a standalone executable file, which can be exported to the client side, and ensures its execution. A user-friendly cross-platform interface is created that performs data check, reports errors and hints at possible ways to correct them.
GUI-BioPASED performs error check of PDB file provided by the user, containing structure description and model parameters, to meet the aminoacid and nucleotide topology library of BioPASED program. PDB file is uploaded to the server to analyze. If it has critical errors, a minute description is shown. If errors are insignificant, the program corrects them and offers the correct PDB file for download. Command file creation for an in silico experiment is automated as much as possible, that almost excludes task formation errors even for beginners. BioPASED program supports both Microsoft Windows and Linux operating systems. Depending in the oprating system installed the user is suggested to download an archive with a proper version of the program. GUI-BioPASED program supports a variety of web browsers: Microsoft Internet Explorer (version 6.0 or greater) and all web browsers based on its core (for example, Maxthon), Mozilla FireFox (version 2 and 3, including Linux builds), Opera, Google Chrome. Some minor differences in page layout and look and feel of controls may occur, but they do not affect the functionality however.
GUI-BioPASED program interface supports English and Russian languages. Along with the client side, GUI-BioPASED program includes an administration module that allows to set up different aspects of functionality adapting it to possible new features that BioPASED may suggest.
If you use the BioPASED, or GUI-BioPASED, or both programs in any work distributed or published, please include the following reference:
Popov A.V., Vorob'ev Yu.N. 2010. GUI-BioPASED program for molecular dynamics modelling of biopolymers with a graphical user interface. Molecular Biology (Moscow). 44. 735–742.
Links on the topic: