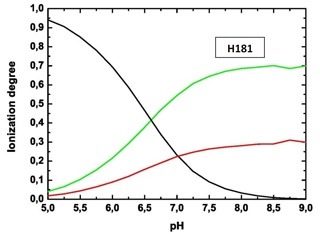
Titration curve of His-181 residue in 1E1A protein.
Histidine residue has unique properties because of his рК0 equal to 6.6, i.e. not far from neutral pH value ~7.0 in physiological conditions. Besides the neutral histidine residue exists in two tautomeric forms, whereas the charged residue has the only chemical structure. Thus the histidine residue in proteins is able to change its chemical structure of the neutral form depending on local environment and energetic preference for either ionized state or neutral state tautomers. Besides neutral aminoacid carboxyl group also exists in two tautomeric forms, with a hydrogen atom at either oxygen atom. Probability for different neutral acid group structure is also determined by the energy which in turn depends on local atomic structure.
The developed method of simultaneous equivalent accouting for mulitple chemical structures of neutral forms of histidine and aminoacid carboxyl groups requires lots of electrostatic energy calculations for a water solvated protein in different ionization states. Such calculations are performed using an optimized fambe_pK_mH algorithm (Fast Adaptive Multigrid Boundary Element). The latter is actually a generalized version of fambe_pH method developed earlier in conformity with a worldwide accepted two state model, namely with charged and common neutral states. The fambe_pK_mH method summarizes the two state model to a multiple neutral states model.
Extending a number of different neutral state atomic structures greatly improves reliability of estimation of ionization state ratio of aminoacid residues, along with different tautomeric fractions of carboxyl groups and neutral histidine residues.
If you use the FAMBE-pK program in any work distributed or published, please include the following reference:
Popov A.V., Vorob'ev Yu.N. 2010. GUI-BioPASED program for molecular dynamics modelling of biopolymers with a graphical user interface. Molecular Biology (Moscow). 44. 735–742.
Links on the topic: